Claudia Solís-Lemus receives NSF CAREER Award

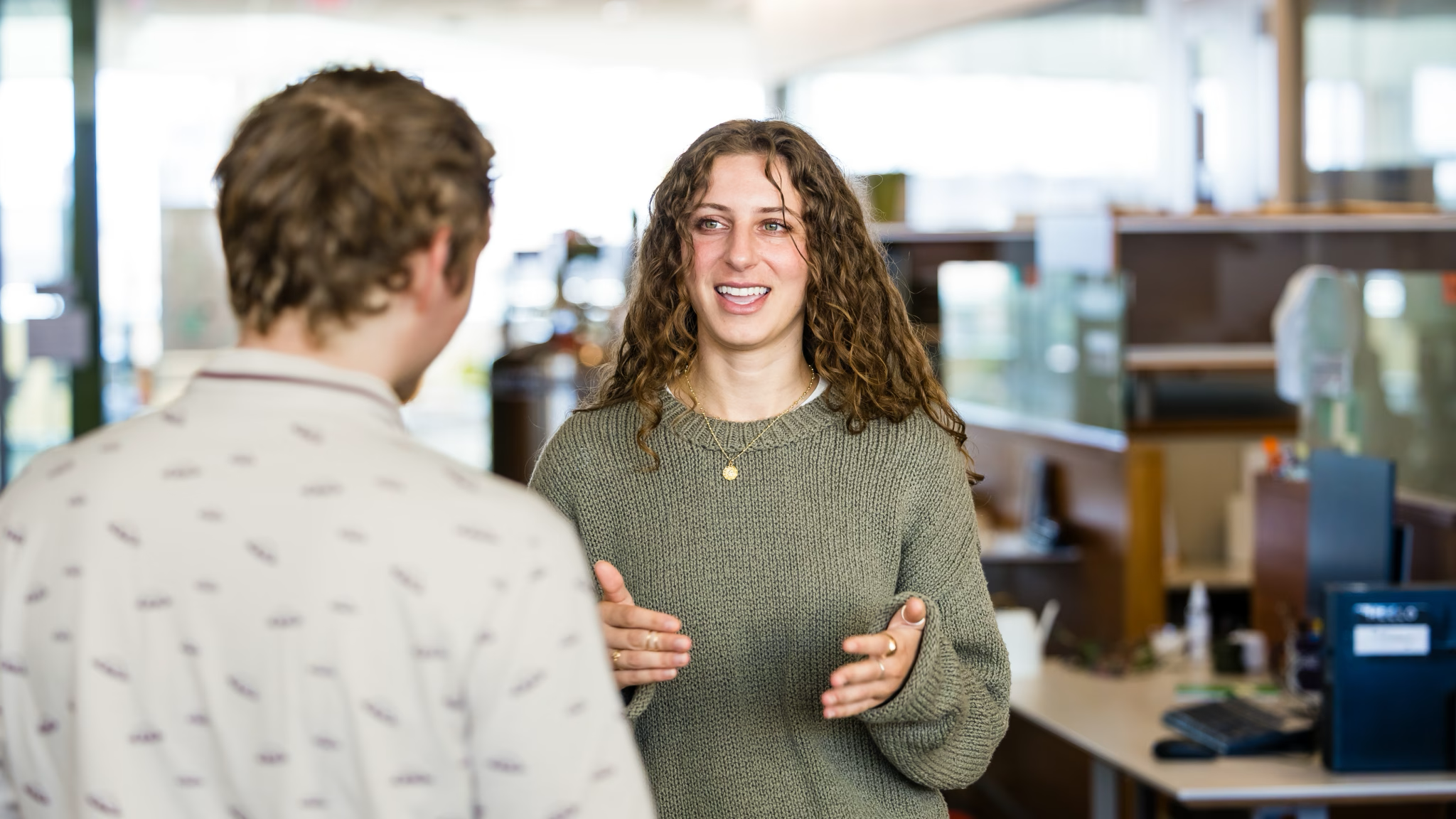
Claudia Solís-Lemus
Claudia Solís-Lemus’ curiosity about the Tree of Life has led to a coveted five-year research grant from the National Science Foundation’s Faculty Early Career Development (CAREER) Award. Each year, this esteemed award provides support for around 50 early-career faculty across the U.S. who show promise as future leaders in science. Solís-Lemus’ NSF grant will support her research, which combines statistical theory and biology to help understand how the biodiversity that we see on Earth evolved from single-cell organisms.
A visual representation of the sheer variety of life evolved from one single celled organism is often depicted as a tree. This Tree of Life helps us understand how biological traits develop through climate change, environmental influences, and the evolution of pathogens. Ultimately, developing tools to reconstruct the tree of life could assist with human health research.
Solís-Lemus’ research proposal focuses on building computational tools that will allow biologists to input their DNA sequences and get a network representation of the evolutionary relationships among their organisms of interest. Her phylogenetic network methodologies will adapt and expand beyond current computational restrictions of big biological data.
NSF CAREER awards require both research and educational outreach components. Solís-Lemus is recognized as an Assistant Professor in the department of Plant Pathology and a faculty member at the Wisconsin Institute for Discovery who exemplifies the role of a teacher-scholar. Her innovative ideas about merging statistics with biology are leading to advances in big data scalability as well as drawing in a younger generation of statisticians.
Solis-Lemus is collaborating with the WI Fast Plants project, first developed at the University of Wisconsin – Madison. FastPlants grow incredibly quickly, featuring seeds that complete a life cycle in about a month. She focuses on middle school biology classes that use FastPlants and has added a data science component. Anyone can log in their collected data from the plants and can do data analysis.
“Our intention is for the students to learn data science along with biology and spark interest in the field of statistics at an early age,” commented Solís-Lemus. “I want to show that statistics is a tool that is useful and is not daunting or scary.”
Additionally, Solís-Lemus is excited to contribute statistical knowledge to phylogenetic research. Solís-Lemus will use an extension of an existing algorithm she developed as a student, building on probability models and extending them to reconstruct larger and more complex networks. Currently, most existing network methods can only estimate networks of a handful of species at most. Solís-Lemus wants to tackle hundreds or even thousands of species. With significant assistance from the CHTC Center for High Throughput Computing and a small server, she will scale and process big data by accounting for hybridization and include a guarantee of identifiability. Allowing the type of parameter values to be distinguished and the hybridization effect on network analysis is important to tackle the complexity of the evolution networks in fungi, viruses or prokaryotes such as bacteria. The methods will unify organisms that currently do not follow the Tree of Life and make space for them. It will show a bigger picture of evolution across all aspects of life.
Claudia eventually wants her algorithms, with their scalable deductive procedures, to be free and open to the public. The application possibilities are endless in regard to evolutionary biology, systematics, conservation and human health. Her quest to identify complex phylogenetic networks could yield important information on detangling of the networks in the Tree of Life.
–Laura C. RedEagle