
Peter Lewis became interested in gene expression when he was in high school after reading a Scientific American article by Robert Tjian, a biochemist who specializes in gene transcription.
“Tjian was talking about gene expression and how that works and all these various pathways. I was really into electric engineering at the time, and I thought it was like electrical circuitry but in a biological context,” Lewis says. Lewis eventually got to know Tjian some years later, when he was a graduate student at the University of California, Berkeley.
When he was an undergraduate in chemistry and biology at the University of Virginia and later as a post-doc at Rockefeller University, David Allis – who Lewis dubs the “Godfather” of epigenetics – was a major influence.
“This was in the late 1990s when histone modifications were just being identified as being causes of gene activation or gene modification. Sometimes at joint floor lab meetings, Allis’s lab would get up and start talking about a new histone modification,” Lewis says. “It was fun to watch all of this happen.”
Coincidentally, epigenetics grew by leaps and bounds as the Human Genome Project unfolded in the 1990s and the early 2000s.
“I was really into electric engineering at the time, and I thought it was like electrical circuitry but in a biological context.”
-Peter Lewis
Despite the success of the Human Genome Project in sequencing the genome, Lewis recognizes that some of its initial promise was not realized and hopes that epigenetic research will supply some missing links. He explains that genome sequencing in-and-of itself once held out the promise of understanding the origins of disease.
“The logic behind the Human Genome Project is that human genes determine disease or aging, and that they might be used for therapy,” Lewis says. “But when we sequenced the human genome, we found that there were only about 21,000 genes. In the past 15 years, we’ve identified what we call ‘missing’ heritability. This means we can’t explain a large number of diseases just based on looking at genes. That was kind of a letdown.”
Though traditional Mendelian genetics cast DNA into a starring role to explain transgenerational inheritance, epigenetic scientists like Peter Lewis seek to discover new mechanisms of inheritance that occur outside of changes to the DNA sequence.
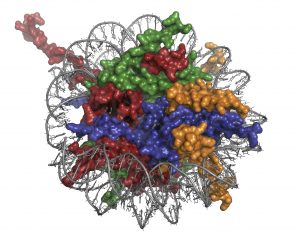
In the cell nucleus, DNA is deeply intertwined with different proteins known as histones. Amazingly, these proteins compress six feet of DNA into a nucleus that is only 3-10 microns in diameter. In weight, chromosomes are 50 percent DNA and 50 percent protein, the most abundant of which are histone proteins. Chromatin, the complex of genomic DNA and histones, makes up chromosomes in cells.
Epigenetics is the study of inheritance that occurs outside of DNA sequence, and chemical modifications to these histones are implicated as one mechanism that affects the genome and the life of the cell. Capable of switching genes “on” or “off,” changes to these histones are also likely caused by environmental factors such as lifestyle and diet.
Only a small percentage of our 21,000 or so genes are “turned on” or expressed in each of the dozens of different cell types found throughout the body. It is now believed that aberrant histone modifications may play a critical role in triggering diseases, such as obesity and cancer, because of the way they cause certain genes to turn on or off. Lewis seeks to discover more about how alterations to the normal pattern of histone modifications can lead cells to divide fecklessly and grow into cancerous tumors.
Lewis, assistant professor of biomolecular chemistry, joined WID in September 2013 where he runs one of four epigenetics labs. By the end of 2014, Lewis’s WID team, in collaboration with some of Lewis’s former colleagues from Rockefeller University and McGill University, discovered the biochemical mechanism of how a mutated histone drives a rare type of childhood bone cancer called chondroblastoma. Lewis is hopeful that this research on chondroblastoma will result in a pharmaceutical therapy that would short circuit tumors driven by the oncogenic histone protein, or oncohistone as he calls it.
This work builds on research Lewis had conducted as a postdoc at Rockefeller University on understanding how an oncohistone drives a fatal pediatric brain cancer known as DIPG (Diffuse Intrinsic Pontine Glioma).
“When I was a postdoc, a paper came out of a group which asked, ‘what are the mutations that give rise to DIPG?’ And they found a highly recurrent mutation in a histone gene. This was the first time that histones were found to be mutated in any kind of disease,” Lewis says. Lewis immediately set to work finding collaborators at Memorial Sloan Kettering Cancer Center.
Lewis’s papers on DIPG research were published in Science in 2013 and 2014.
“Eventually, we figured out the biochemical mechanism of how that works, and then we began to model it. We made a lot of progress understanding DIPG,” he says.
His research team addressed questions like: What is the cell of origin that gives rise to the tumor? What are the developmental pathways that are misregulated? How does the oncohistone mutation block normal developmental pathways?
“If you begin to work backwards in understanding how the tumor arose in the first place, you might be able to figure out what pathways the tumor still needs to grow, and if you were somehow to perturb that mechanism, then a therapy could be devised,” Lewis says.
In addition to studying the mechanism of oncohistones, the lab also studies the mechanism of a chaperone protein that assists with putting histones onto the DNA—a process that seems to insure the health of the genome. It has been recently discovered that mutated Daxx, and its partner in crime, ATRX, are present in 60 percent of a rare type of pancreatic cancer. During his postdoc, Lewis and colleagues discovered that the ATRX-Daxx protein complex functions as a histone chaperone.
Once again, Lewis is hoping to answer some basic science questions about Daxx.
Lewis is asking: How does Daxx function as a histone chaperone to deliver the histones to the DNA? And where in the genome does it do this?
“We know that it’s involved in maintaining telomeres, the caps of the chromosomes, which are really important for maintaining genome integrity. We know that Daxx helps put histones in at telomeres, and we think it helps keep those caps nice and tight. At the moment we are working to understand how Daxx functions at other parts of the genome,” Lewis says.
“We’re also examining the tumors. We have some neuroendocrine tumors without Daxx and some with Daxx. How does loss of Daxx change the epigenomic landscape? And how do these changes contribute to the gene expression profile that drives the metastatic potential of these tumors?”
Lewis believes that epigenetics may yield answers to the origins of certain diseases.
“People with exactly the same genotype can have different outcomes depending on their environment, and how the environment affects the biology,” he says.
–Mary Sussman

You must be logged in to post a comment.