Modeling is the process of developing and testing mathematical representations of processes or events.
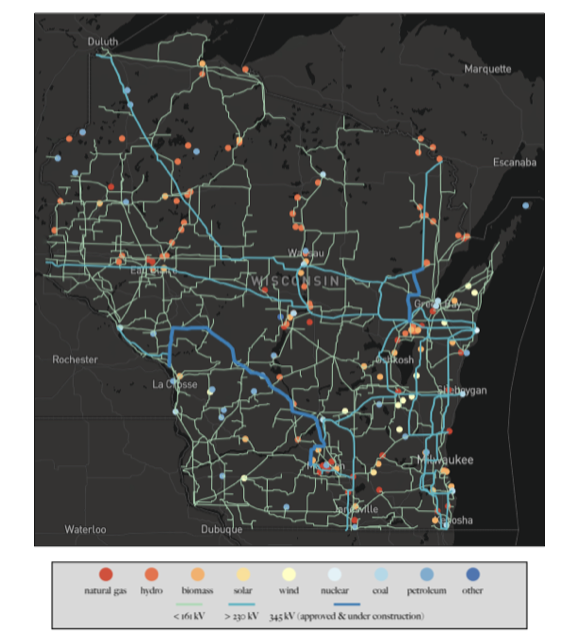
Modeling is foundational to much of the work done at WID, from modeling evolutionary and biological systems to power grids and fish habitats. Researchers at WID work on the fundamental algorithms and formulations that allow them to build the best and most useful models to solve difficult problems.
Modeling is a key tool for WID’s Data Science Hub.